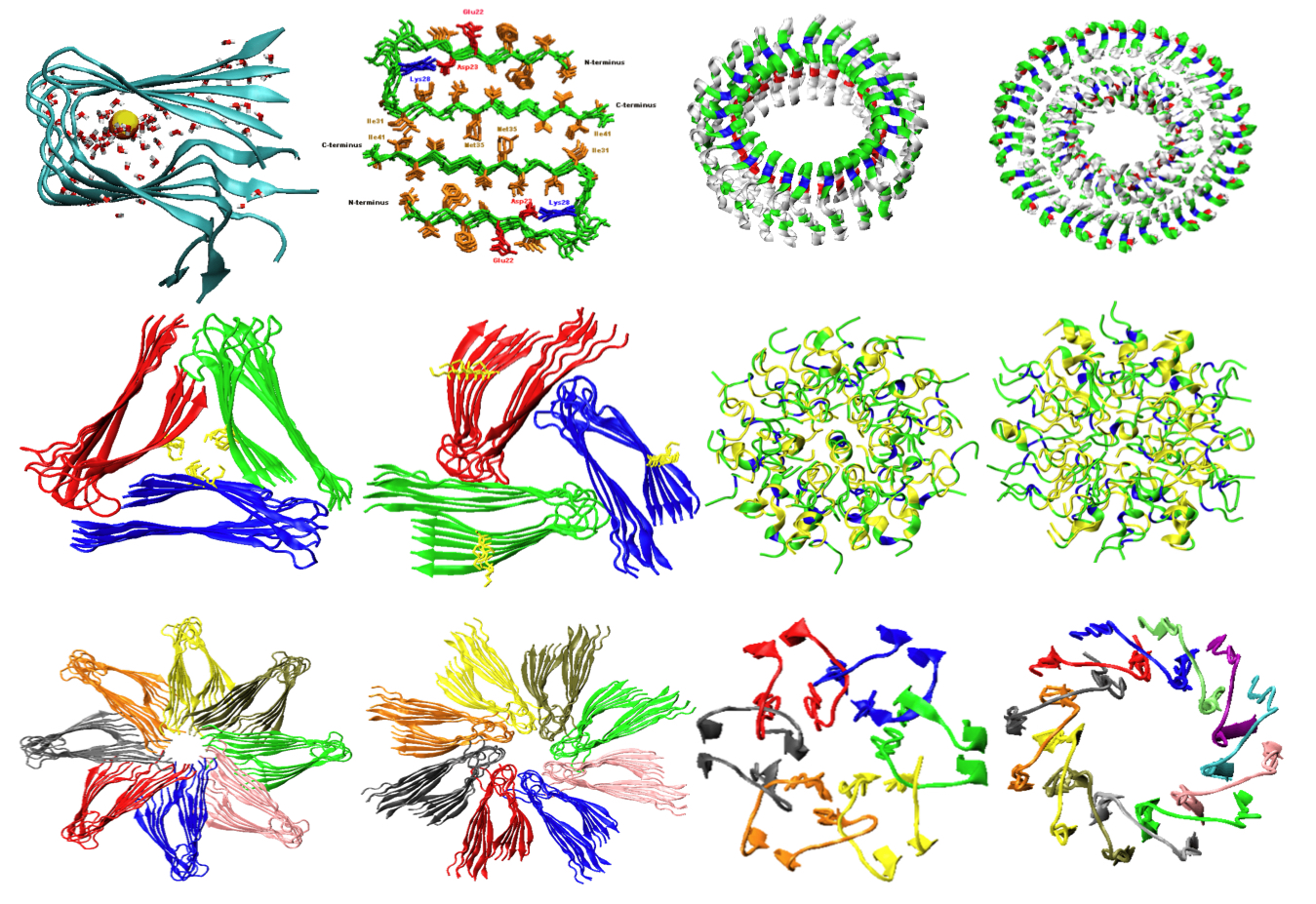
A University of Akron researcher is creating sophisticated computer simulations at the Ohio Supercomputer Center to help understand how “misfolded” proteins in the brain contribute to degenerative disorders, such as Alzheimer’s disease.
Alzheimer’s disease is the most common human neurodegenerative disorder, affecting as many as 5.1 million people in America alone, according to the National Institutes of Health (NIH). Alzheimer’s most often appears after age 60 and leads to progressive and irreversible memory loss, disability and, eventually, death. Disorders like Alzheimer’s occur through a complex series of events that take place in the brain over a long period of time, probably beginning a decade or two before the most significant symptoms arise.
In the nucleus of nearly every human cell, long strands of DNA are packed tightly together to form chromosomes, which contain all the instructions a cell needs to function. To deliver these instructions to various other cellular structures, the chromosomes dispatch very small protein fibers – called oligomers – that fold into three-dimensional shapes. Misfolded proteins – called amyloid fibrils – cannot function properly and tend to accumulate into tangles and clumps of waxy plaque, robbing brain cells of their ability to operate and communicate with each other, according to NIH.
The strength of this research project lies in the integration of different experimental techniques with a computational approach to effectively illustrate various aspects of amyloid formation and its damaging effects, according to Jie Zheng, Ph.D., an assistant professor of chemical and biomolecular engineering at the University of Akron.
“The exact mechanism of amyloid formation and the origin of its toxicity are not fully understood, primarily due to a lack of sufficient atomic-level structural information from traditional experimental approaches, such as X-ray diffraction, cryoelectron microscopy and solid-state NMR data,” Zheng explained. “Molecular simulations, in contrast, allow one to study the three-dimensional structure and its kinetic pathway of amyloid oligomers at full atomic resolution.”
Zheng’s research group is developing a multiscale modeling and simulation platform that integrates structural prediction, computational biology and bioinformatics to establish a direct correlation between the formation of oligomers and their biological activity in cell membranes. This research is important for understanding the build-up of protein plaque, how it contributes to the breakdown of cells and how the process might be prevented.
“This project has broad impacts on the prevention of diseased-related protein misfolding and aggregation, said Zheng. “The ultimate goal of this project is to rationally design a series of effective ligands/inhibitors to prevent amyloid formation.”
Zheng’s anti-amyloid project is leveraging the computational muscle of the IBM Cluster 1350 system at the Ohio Supercomputer Center. The center’s flagship supercomputer system, named the Glenn Cluster, features 9,500 cores, 24 terabytes of memory and a peak computational capability of 75 teraflops – about 75 trillion calculations per second.
Zheng’s computational approach uses replica-exchange molecular dynamics simulations, a computationally intensive process for analyzing protein folding. The REMD method performs a large number of concurrent simulations of amyloid formation while introducing different variables, such as temperature, that can influence the outcome of the process.
“It’s great to hear that OSC resources are being utilized to investigate the cause of such devastating human diseases, especially one that robs us of the very essence of our humanity, our memories,” said Don Stredney, director of the Center’s Interface Lab and research scientist for Biomedical Applications. “We are pleased that OSC can accelerate the iterations required for such complex conformational studies, leading to new insights into the mechanisms of memory loss.”
Zheng’s amyloid protein research project recently won a five-year, $400,000 award through the Faculty Early Career Development program of the National Science Foundation (NSF). One of the NSF’s most prestigious recognitions, the CAREER Award is bestowed “in support of junior faculty who exemplify the role of teacher-scholars through outstanding research, excellent education and the integration of education and research within the context of the mission of their organizations.”
Zheng joined the department of Chemical and Biomolecular Engineering at the University of Akron in 2007 as an assistant professor, after working as a scientist at the National Cancer Institute since 2005. He received his doctorate in chemical engineering from the University of Washington in 2005.
The Ohio Supercomputer Center is a catalytic partner of Ohio universities and industries that provides a reliable high performance computing infrastructure for a diverse statewide/regional community. OSC promotes and stimulates computational research and education in order to act as a key enabler for the state's aspirations in advanced technology, information systems, and advanced industries. For additional information, visit http://www.osc.edu