Researchers at The Ohio State University (OSU) are applying the power of supercomputers to a small plant in the mustard family to better understand how complex genetic processes can lead to different types of cells.
Dan Siegal-Gaskins, a postdoctoral fellow affiliated with the Mathematical Biosciences Institute and the Grotewold Lab in OSU’s department of Plant Cellular and Molecular Biology, is leveraging resources at the Ohio Supercomputer Center (OSC) as part of a larger study of cell differentiation in the model plant Arabidopsis thaliana.
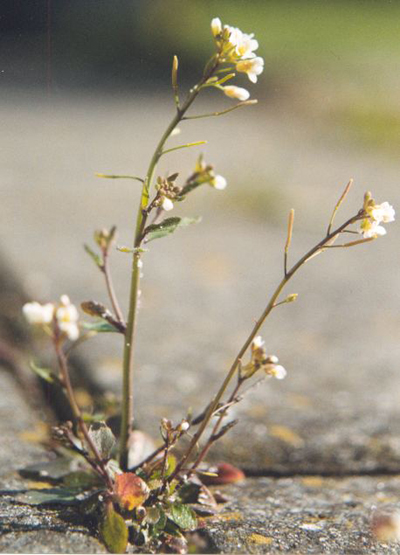
Known commonly as thale cress or mouse-ear cress, Arabidopsis has one of the smallest genomes in the plant kingdom and plays a role in developmental biology similar to that of mice and fruit flies. The small size and short growing period of Arabidopsis makes it particularly well-suited for genetic studies.
At a specific phase of Arabidopsis leaf development, cells on the surface of the leaf receive genetic instructions to become either one of the majority ‘pavement’ cells or a large hair-like cell known as a trichome. The specific function of trichomes is unclear, although they may be involved in preventing infection, protecting delicate tissues on the underside of the leaf, or reducing the amount of water lost to evaporation.
To better understand how cells develop into trichomes, Siegal-Gaskins, colleague Kengo Morohashi and Principal Investigator Erich Grotewold are focusing on relationships between three proteins that figure prominently in determining a cell’s fate. Most importantly, the researchers are supplementing traditional benchwork with mathematics to better understand the proteins’ functional relationships.
The mathematical model Siegal-Gaskins constructed consists of seven differential equations and twelve unknown factors. For his preliminary studies, he turned to OSC to choose random values for the unknowns and solve the equations for millions of different random value sets.
“Due to the large range of possible parameters and the complexity of the problem, we took advantage of OSC’s parallel processing capabilities and the MATLAB computing environment,” Siegal-Gaskins said. “This process was repeated for five million randomly-chosen parameter sets, and the set that gave us the closest agreement with experimentation was kept.”
To meet the challenge of processing the millions of iterations, Siegal-Gaskins accessed OSC’s IBM Cluster 1350. The center’s flagship supercomputer system, nicknamed the Glenn Cluster, features 9,500 cores, 24 terabytes of memory and a peak computational capability of 75 teraflops – which translates to 75 trillion calculations per second.
“Dr. Siegal-Gaskins is leveraging high performance computing (HPC) to better understand biological systems at the cellular and molecular level,” said Yuan Zhang, client and technology support engineer at OSC. “His project is especially well-suited for the Glenn Cluster, which is largely dedicated to research in the biosciences, and MATLAB software, which features many tools for numerical computations.”
The MATLAB programming software package is described as “a technical computing environment for high-performance numeric computation and visualization” that produces output in mathematical formats. OSC has been a leader in running MATLAB and other scripting languages in HPC environments.
“Our bcMPI software, initially released in 2006, interfaces with HPC cluster technologies when executing MATLAB scripts on a cluster,” explained David Hudak, director of HPC engineering at OSC. “Over the last year, we have been working to improve the accessibility of parallel MATLAB. We designed Remote MATLAB Services (RMS) to enable our users to transition MATLAB scripts developed on their laptops to HPC resources. Dan was an early adopter of OSC RMS, and we learned a lot from his feedback. It was a very good fit for his needs.”
With the combination of computational modeling, literature-based analyses and laboratory experimentation, Siegal-Gaskins and Morohashi determined that the three cell fate proteins seem to constitute an “incoherent feed forward loop,” a relationship in which a master regulator (MR) triggers expression of two genes involved in the initiation of trichome cell development, one of which (G1) later suppresses expression of the other (G2).
Siegal-Gaskins described their research approach as typical of the growing field of ‘systems biology’: “In addition to the wide array of tools typically used in the biological sciences, a ‘systems biologist’ uses tools from the physical sciences – physics, engineering, mathematics, and computer science – to probe living systems in an attempt to understand biological function.”
Indeed, revolutionary advances in biological science and technology have challenged scientists to develop new mathematical, statistical and computational methods for analyzing and structuring large amounts of data.
“Mathematical modeling is now used as an integral tool in the study of biological systems, both for the generation of hypotheses and to lend support to experimental results,” Siegal-Gaskins said. “As we attempt to understand how it is that cells can ‘choose’ a fate, modeling can play a particularly crucial role.
The scarcity of experimental data would make it difficult to figure out exactly what is going on inside cells if there wasn’t some mathematics and computational work to back it up and fill in the blanks.”
The Ohio Supercomputer Center is a catalytic partner of Ohio universities and industries that provides a reliable high performance computing infrastructure for a diverse statewide/regional community. OSC promotes and stimulates computational research and education in order to act as a key enabler for the state's aspirations in advanced technology, information systems, and advanced industries. For additional information, visit http://www.osc.edu