“The origins or causes of diseases determine how they manifest illness and what we can do about them,” explained Matthew T. Weirauch, Ph.D., an associate professor in the Center for Autoimmune Genomics and Etiology at Cincinnati Children’s Hospital and the pediatrics department at the University of Cincinnati College of Medicine. “Of the known genetic risk variants for diseases, about 90 percent likely alter gene regulation, but almost nothing is known about how they do this.”
Weirauch’s research team at Cincinnati Children’s is studying “how they do this” by analyzing how these genes alter disease risk and interact with the environment. With that knowledge, he believes, scientists might be able to leverage these mechanisms to develop specific therapies and preventive measures for a variety of conditions, such as multiple sclerosis, rheumatoid arthritis, type 1 diabetes, inflammatory bowel disease and lupus.
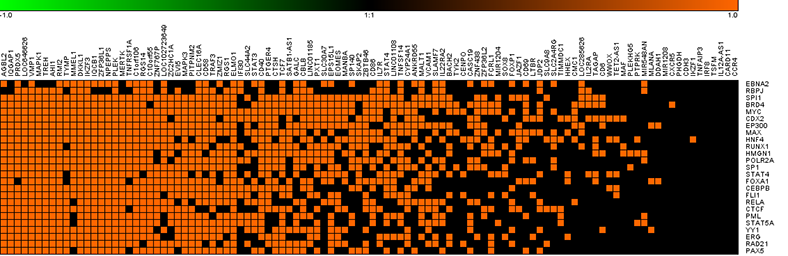
Using high performance computing and storage systems at the Ohio Supercomputer Center, the team created the Regulatory Element Locus Intersector (RELI) algorithm for estimating the significance of the overlap between disease-associated variants and large collections of transcription factor (TF) genome-wide binding datasets. TFs are proteins that control when and where in the body genes get “turned on” and “turned off.” In this regard, they are key players in translating the DNA-encoded genome into RNA, which then creates the proteins that make cells (and life) work.
The team has discovered 2,264 associations of TF binding sites with multiple disease-associated genome regions related to 93 human disease or physical conditions. Its findings potentially identify molecular participants in the regulatory mechanisms that initiate disease, their contributing genetic mutations and the cell types involved in the developmental pathways of disease.
Prior to the advent of highthroughput genotyping and sequencing, it was widely assumed that an understanding of the cause of a disease could be achieved by studying one genetic variant at a time. It is now appreciated that most diseases are generally a manifestation of the additive action of many variants tipping the scales towards increased disease risk.
“We have created an innovative computational method that leverages the wide availability of data describing TF binding interactions with the genome to deliver a solution analyzing multiple genetic locations simultaneously,” said Weirauch. “We expect that this project represents an important initial step on a long journey exploring the association between the binding of TFs and the onset of many human diseases.”
_________
PROJECT LEAD // Matthew Weirauch, Ph.D., Cincinnati Children's Hospital Medical Center
RESEARCH TITLE // Gene regulation in disease genetics
FUNDING SOURCES // National Institutes of Health, Alliance for Lupus Research, Cincinnati Children’s Hospital Research Fund
WEBSITE // cincinnatichildrens.org/bio/w/ matthew-weirauch